Methods
The cryo-EM method has become very powerful in the past several years and for many structural biologists it is becoming the method of choice. The full potential of the technique, however, is still to be realized as the scientific community is in great need of hardware-based and software improvements. Although the first 3D images of biological specimens at true atomic resolution have been obtained recently, the routine resolution of cryo-EM still lags behind in detail and quality in comparison with common material science electron microscopy as it is imaged at the Ernst-Ruska Centre (ER-C) 1 and 2. Therefore, in collaboration with the ER-C-1 and ER-C-2 we will apply new imaging hardware to biological samples. Our technique development also includes novel sample preparation, data acquisition and image processing methods that we benchmark using biological test specimens. We apply these innovative cryo-EM methods to the structures of challenging biological systems with a particular focus on membrane assemblies.
Publications
- Weis F., Beckers M., von der Hocht I., Sachse C. (2019) Elucidation of viral disassembly switch of tobacco mosaic virus. EMBO Reports 20(11):e48451.
- Fromm, S.A., Bharat, T.A.M., Jakobi, A.J., Hagen, W.J.H., and Sachse, C. (2015). Seeing tobacco mosaic virus through direct electron detectors. J Struct Biol 189, 87–97.

Software
SPRING
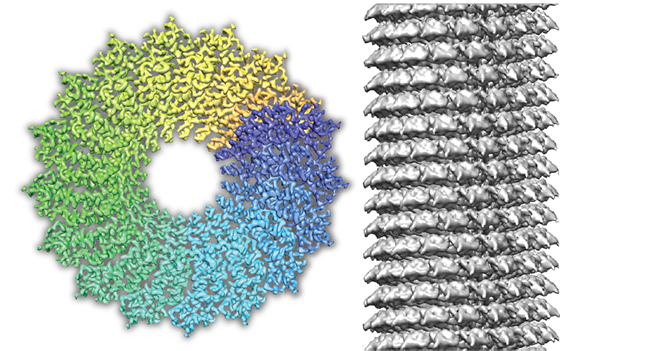
SPRING is a single-particle based helical reconstruction package developed in our laboratory to determine 3D structures of a variety of highly ordered and less ordered specimens with helical symmetry from electron micrographs.
- Desfosses, A., Ciuffa, R., Gutsche, I., and Sachse, C. (2014). SPRING – an image processing package for single-particle based helical reconstruction from electron cryomicrographs. J Struct Biol 185, 15–26.
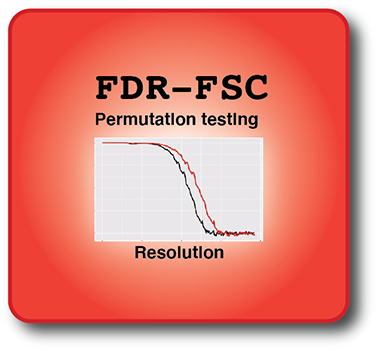

FDR-FSC
FDR-FSC is a robust cryo-EM map resolution estimation tool that thresholds the Fourier Shell correlation curve by statistical permutation testing including false discovery rate control. The procedure works unsupervised as it accepts untreated half-maps and does not require any solvent-flattening masking.
- Beckers M., and Sachse C. Permutation testing of Fourier shell correlation for resolution estimation of cryo-EM maps. (2020) J Struct Biol:107579.
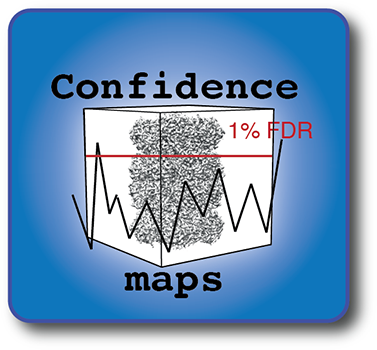

Confidence maps
FDRthresholding is a density interpretation tool to analyze cryo-EM structures using a common objective threshold of “false discovery rate” with respect to background noise. It is executable as part of the CCPEM suite and the SPOC package.
- Beckers, M., Jakobi, A. J., Sachse, C. (2019) Thresholding of cryo-EM density maps by false discovery rate control. IUCr Journal 6 (1).
- Beckers M., Palmer C.M., and Sachse C. Confidence maps: statistical inference of cryo-EM maps. (2020) Acta Crystallogr Sect D Struct Biol 76(4):1–8.
LocScale
LocScale is density scaling (sharpening) procedure that aims to enhance interpretability of cryo-EM density maps. LocScale is a reference-based local amplitude scaling tool using prior model information to improve contrast of cryo-EM density maps. It is executable as part of the CCPEM suite.
- Jakobi, A. J., Wilmanns, M., and Sachse, C. (2017). Model-based local density sharpening of cryo-EM maps. eLife; 6:e27131.
